Eukaryotic cells control the ultimate outcome of their genetic programs not only by regulating the synthesis and processing of transcripts but also by controlling their export, localization, translatability, and stability. We discovered that the transcription apparatus mediates the association of specific RNA-binding proteins with nascent transcripts. These proteins subsequently accompany the mRNA targets to the cytoplasm and “imprint” their future behavior, we refer to them as RNA imprinting.
Few dozens of imprinting proteins were identified, using our PROFIT methodology
mRNA imprinting is widespread, mediated by the Pol II subunit Rpb4 and is responsive to environmental cues. We identified several unexpected imprinting proteins, including translation factors, protein chaperons, transcription factors and mRNA decay factors.
Outstanding imprinting factors reported in Urim et al.
o Translation factors: eIF4G eIF4E and eIF3 component Rpg1 bind the nascent Pol II transcripts co-transcriptionally and subsequently function in translation.
o mRNA decay factors: Xrn1 and Rat1.
o Protein chaperones: Hsp70 variants Ssa1 and Ssa2.
o Transcription factors: Taf14, Sfp1 and Spt6. Spt6, a component of the FACT complex acting during transcription elongation, was previously shown to regulate mRNA stability in the cytoplasm.
o Substrate-delivery proteins of the proteasome and mitochondrial-targeting factors.
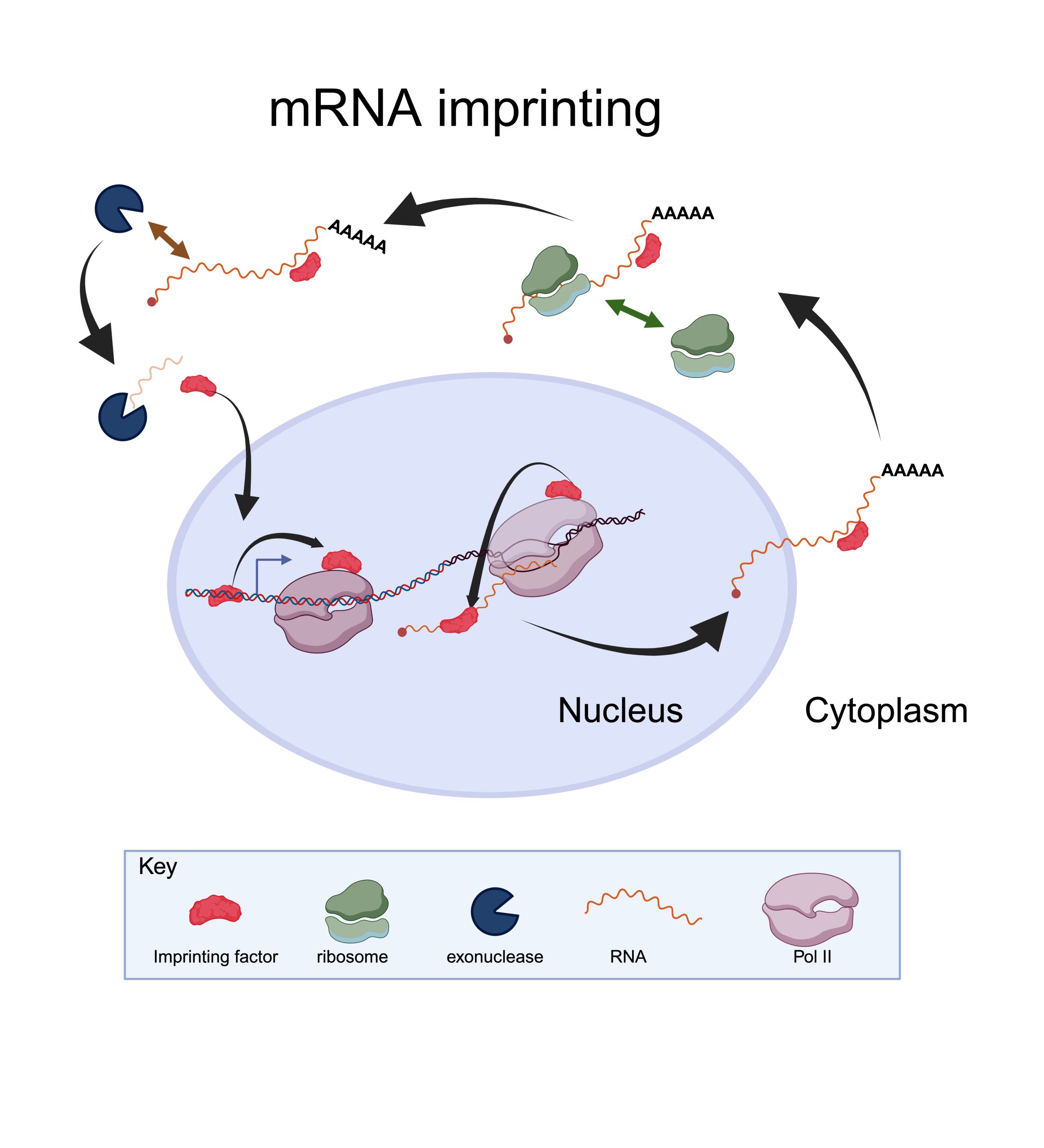

Model of mRNA imprinting, based on Kelbert et al., 2024. The imprinting factor binds to
specific gene promoters, regulating transcription. Upon transcription initiation, it dissociates from
the promoter and associates with the elongation complex by binding to the Pol II subunit Rpb4. When a transcript containing a specific motif emerges from Pol II, the factor binds the nascent RNA motif while dissociating from Pol II. The mRNA/factor complex is subsequently exported to the cytoplasm. The factor then influences the mRNA translatability and stability. After mRNA degradation, the factor returns to the nucleus to start a new cycle.
The model has been evolved in our lab since 2010. See publications:
- Choder, M. (2011). mRNA imprinting: additional level in the regulation of gene expression. Cellular Logistics 1, 37-40.

- Goler-Baron et al. (2008). Transcription in the nucleus and mRNA decay in the cytoplasm are coupled processes. Genes & Dev.22, 2022-2027.

- Harel-Sharvit et al. (2010). RNA polymerase II subunits link transcription and mRNA decay to translation. Cell 143, 552-563.

- Bregman et al. Promoter elements regulate cytoplasmic mRNA decay. (2011). Cell, 147, 1473-83.

- Chattopadhyay et al. RNA-controlled Nucleocytoplasmic shuttling of the exoribonuclease Xrn1 is required for mRNA synthesis and decay and for responses to environmental changes. Nature communication 13(1), 7184-7201. doi: 10.1038/s41467-022-34417-z.
- Kelbert et al. (2024). The transcription factor Sfp1 imprints specific classes of mRNAs and links their synthesis and cytoplasmic decay. eLife 12, RP90766 PMC11446548 https://doi.org/10.7554/eLife.90766.1
- Urim et al. mRNA Imprinting: transcription apparatus can remotely control cytoplasmic post-transcriptional mechanisms by dozens of proteins. eLife, in press.





